Homology and Model Organisms
Introduction
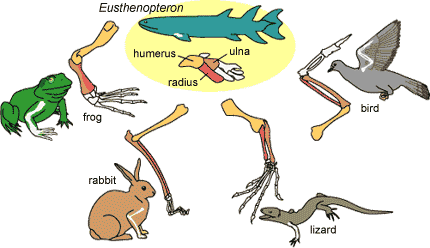
Homology is the presence of similar characteristics shared by organisms due to distant relatedness. This comes from evolutionary theory which dictates that these similar traits are derived from a trait passed down by a common ancestor [1]. We can use homology to study genetics by finding conserved regions within a gene of interest and searching for these similar regions within these homologous genes. By using NCBI BLAST, and HomoloGene, we can find homologs of the PMP22 gene in other organisms [2] [3]. In particular, these sites use the amino acid sequence that comprise the gene in order to compare it to other organisms.
Of particular interest to us is finding homologs within organisms that are considered model organisms. Model organism such as mice are excellent for research as they are well studied and their genomes are well understood already, so by researching a particular gene in a model organism, the information can be understood and hopefully transferred back into the understanding of humans [4].
Of particular interest to us is finding homologs within organisms that are considered model organisms. Model organism such as mice are excellent for research as they are well studied and their genomes are well understood already, so by researching a particular gene in a model organism, the information can be understood and hopefully transferred back into the understanding of humans [4].
Results
|
|
|
|
Discussion
PMP22 has homologs in many model organisms, including the ones shown above. All of the organisms which possess a PMP22 homolog, also have an extensive peripheral nervous system, which makes sense since PMP22 is known to be highly involved in the formation of myelin in the peripheral nervous system in human. Having this knowledge of homologs of PMP22 allows us to select a model organism for which PMP22 can be studied in.
References
[1] Sommer, R. J. (2008). Homology and the hierarchy of biological systems. BioEssays, 30(7), 653-658. doi:10.1002/bies.20776
[2]Blast: Basic local alignment search tool. (n.d.). Retrieved May 07, 2021, from https://blast.ncbi.nlm.nih.gov/Blast.cgi
[3]Home - homologene - ncbi. (n.d.). Retrieved May 07, 2021, from https://www.ncbi.nlm.nih.gov/homologene
[4]Research organisms. (n.d.). Retrieved May 07, 2021, from https://www.nigms.nih.gov/education/fact-sheets/Pages/using-research-organisms.aspx
[5] Homology example picture - Homologies. (n.d.). Retrieved May 07, 2021, from https://evolution.berkeley.edu/evolibrary/article/0_0_0/lines_04
[2]Blast: Basic local alignment search tool. (n.d.). Retrieved May 07, 2021, from https://blast.ncbi.nlm.nih.gov/Blast.cgi
[3]Home - homologene - ncbi. (n.d.). Retrieved May 07, 2021, from https://www.ncbi.nlm.nih.gov/homologene
[4]Research organisms. (n.d.). Retrieved May 07, 2021, from https://www.nigms.nih.gov/education/fact-sheets/Pages/using-research-organisms.aspx
[5] Homology example picture - Homologies. (n.d.). Retrieved May 07, 2021, from https://evolution.berkeley.edu/evolibrary/article/0_0_0/lines_04